|
A mirror website is available at BioEdit site. More tech-savvy users employ more advance graphical software like Photoshops or AutoCAD to create figures, but this is hardly ideal.įinally, as of 2021, the original BioEdit website host at NCSU is no longer available, as its maintainer Dr. However, its free version has some limitations, particularly in editing the sequences, forcing the users to use workarounds such as manually editing the sequence in a text editor. Other softwares provide workarounds to generate figures suitable for modern articles. Furthermore, the software is only provided for Windows operating system, thus making it more difficult to run on Mac/Linux environment. The figures generated by BioEdit are also outdated compared to contemporary standards. It has an outdated graphical user interface and is likely to contain unresolved bugs and security compromises. Source 2Īlthough BioEdit is a very popular program, and still widely used as of 2021, its development had been stopped since 2007. It also has the capability to generate figures useful for scientific publications.įigure: Example of figure generated by BioEdit. Several sequence manipulation and analysis options and links to external analysis programs facilitate a working environment that allows the user to view and manipulate sequences with simple point-and-click operations. 
It has a multiple document interface with convenient features that makes the alignment and manipulation of sequences relatively easy on a desktop computer. BackgroundīioEdit 1 is a popular biological sequence alignment editor written originally for Windows 95/98/NT/2000/XP. However, BioEdit is no longer being maintained, and thus we propose the creation of a new open-source software package that aims to replace BioEdit. One of the most widely used tools in use for such tasks is BioEdit 1. You can learn more about sequence alignments on the UniProt help page.Abstract: Sequence management, alignment editing and figure generation is an important task performed daily by biologists. You can also run Alignment from within the Basket. All relevant results pages (such as UniProtKB, UniRef, UniParc and tool results) provide an ‘Align’ button to run alignments directly by selecting entries with checkboxes. The following kinds of UniProt identifiers are supported: P00750Įach UniProtKB entry which contains both a sequence and one or more isoforms of that sequence, enables you to align the canonical sequence and its isoforms. Note – advanced users are given the option of varying the alignment parameters from those given as default.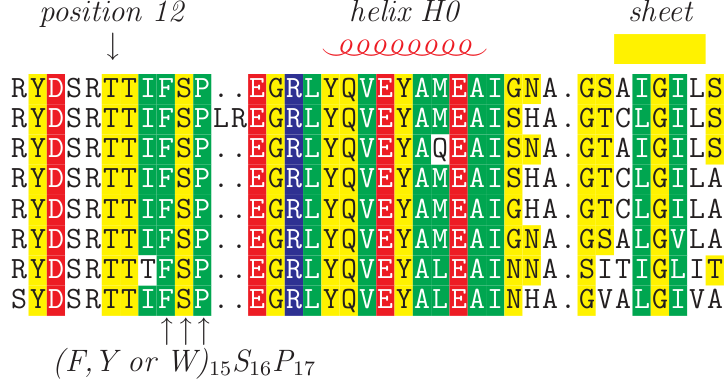 
Enter either protein sequences in FASTA format or UniProt identifiers (as above) into the form field.Click on the Align link in the header bar to align two or more protein sequences with the Clustal Omega program.Exercise: mapping other database identifiers to UniProtĪll materials are free cultural works licensed under a Creative CommonsĪttribution 4.0 International (CC BY 4.0) license, except where further licensing details are provided.Ī sequence alignment is a way of arranging the primary sequences of a protein to identify regions of similarity that may be a consequence of functional, structural, or evolutionary relationships between the sequences.Exercise: finding entries with 3D structures.Downloading a proteome set for specific organism.Accessing UniProt data programmatically.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
